Gabriel Foley
Email: gabriel.foley@uqconnect.edu.au
LinkedIn: gabrielfoley
I'm currently pursuing my PhD under the supervision of Associate Professor Mikael Bodén and Professor Elizabeth Gillam at the School of Chemistry and Molecular Biosciences within the University of Queensland (UQ). I previously completed a Bachelor of Biotechnology (Honours Class I) majoring in Bioinformatics and Innovation Management at UQ.
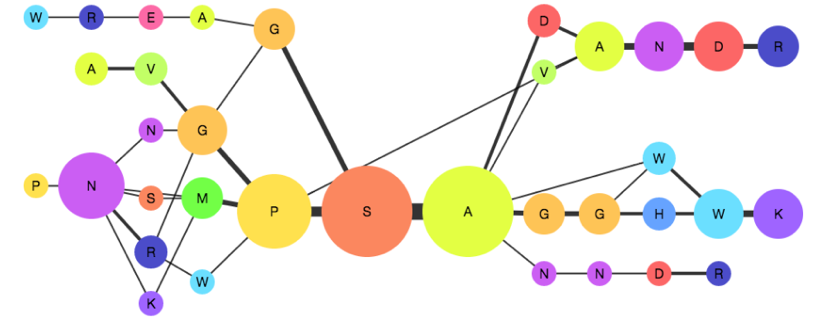
My PhD is focused on exploring the application of probabilistic graphical models—specifically the Partial Order Alignment Graph (POAG) to a number of areas within bioinformatics. I'm currently working on using POAGs as an alternative data structure for multiple sequence alignment to improve error handling within protein alignment. I'm also interested in ancestral sequence reconstruction and how POAGs can serve as a generative model which can be sampled from in order to produce and rank potential ancestors of protein sequences.
I am in the process of implementing a JavaScript application which allows protein engineers to perform partial order alignment and visualise and interact with partial order graphs.
Publications
Williams SJ, Yin L, Foley G, Casey LW, Outram MA, Ericsson DJ, Lu J, Boden, M, Dry IB and Kobe, B (2016). Structure and Function of the TIR Domain from the Grape NLR Protein RPV1. Frontiers in Plant Science, 7. DOI: 10.3389/fpls.2016.01850
Awards
2016 UQ Association of Postgraduate Students Pitch My Research Finalist
2015 Bing and Ross Barnard Biotechnology Prize
2015 SCMB Honours Scholarship
Figure 1: Partial Order Alignment Graph.
Authors
Created by gabe (Gabriel Foley) on 2017/04/20 20:15.
Contributing authors: