This is an old revision of the document!
Evolu-sec package
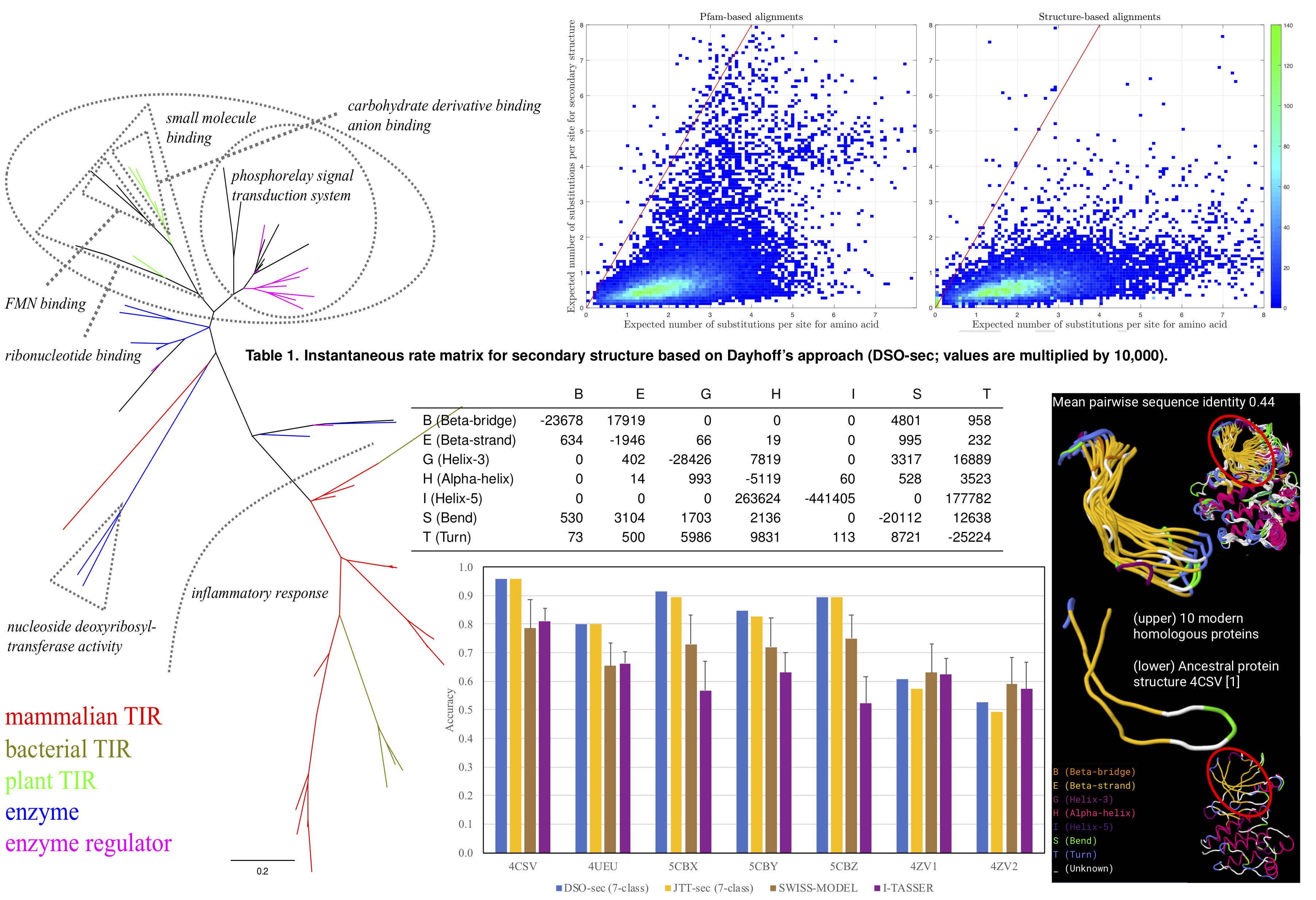
<html> <p class=MsoNormal style='text-align:justify;text-justify:inter-ideograph'><span class=SpellE><span lang=EN-GB>Evolu</span></span><span lang=EN-GB>-sec package forms part of a research project, reported in “Modelling the evolution of protein secondary structure: a new phylogenetic metric suitable for assessing structural similarity”.<o:p></o:p></span></p> <p class=MsoNormal style='text-align:justify;text-justify:inter-ideograph'><span class=SpellE><span lang=EN-GB>Evolu</span></span><span lang=EN-GB>-sec package contains phylogenetic analysis tools based on an evolutionary model of secondary structure <b style='mso-bidi-font-weight:normal'>[centre, Table 1]</b>. The analysis focuses on phylogenetic tree inference <b style='mso-bidi-font-weight: normal'>[left]</b>, evolutionary distance estimation <b style='mso-bidi-font-weight: normal'>[top]</b>, and ancestral secondary structure reconstruction. <b style='mso-bidi-font-weight:normal'>[bottom and right]</b>.</span></p> </html>
The research project abstract is provided in the next section. The dataset for the model development and the software can be accessed in the “Supplementary Data” section.
Research Project
Modelling the evolution of protein secondary structure: a new phylogenetic metric suitable for assessing structural similarity
Jhih-Siang Lai, Burkhard Rost
, Bostjan Kobe and Mikael Bodén
School of Chemistry and Molecular Biosciences, The University of Queensland
Contact: Jhih-Siang Lai
Abstract
<html>
<p class=MsoNormal style='text-align:justify;text-justify:inter-ideograph'><a
name="_GoBack"></a><span lang=EN-GB>Ancestral sequence reconstruction has had
recent success in decoding the origins and the determinants of complex protein
functions. However, attempts to reconstruct ancient proteins and phylogenetic
analyses of remote homologues must handle extreme amino-acid sequence diversity
resulting from extended periods of evolutionary change [2]. We exploited the wealth
of protein structures in the Protein Data Bank (PDB) to develop an evolutionary
model based on protein secondary structure. The approach follows the
differences between discrete secondary structure states observed in modern
proteins and those hypothesised in their immediate ancestors [1]. Based on this new
evolutionary model, we implemented maximum likelihood-based phylogenetic
inference tools to reconstruct ancestral secondary structure. The predictive
accuracy from the use of the evolutionary model surpasses that of structure
comparative modelling and sequence-based prediction methods; the reconstruction
extracts information not available from modern structures or the ancestral
protein sequences alone [3,4,5]. Based on a phylogenetic analysis of a sequence-diverse
protein family, we showed that the model has the capacity to highlight
relationships that are evolutionarily rooted in structure and not evident in
sequence-based phylogenetic analysis.</span></p>
</html>
Supplementary Data
Supplementary data for "Modelling the evolution of protein secondary structure: a new phylogenetic metric suitable for assessing structural similarity" can be downloaded from
Dataset or Software, separately.
Package may be downloaded as a single archive from
Full package download.
- (Ancestral secondary structure reconstruction) ASSR needs version MATLAB 8.2 (R2013b) or later.
- Please send feedback and questions to author.
References
<html>
<body bgcolor=white lang=ZH-TW link=blue vlink="#954F72" style='tab-interval: 24.0pt;text-justify-trim:punctuation'>
<div class=WordSection1 style='layout-grid:20.0pt'>
<p class=MsoListParagraph style='margin-left:18.0pt;mso-para-margin-left:0gd; text-align:justify;text-justify:inter-ideograph;text-indent:-18.0pt;mso-list: l0 level1 lfo1'><![if !supportLists]><span lang=EN-US style='font-size:10.0pt; font-family:"Helvetica",sans-serif;mso-fareast-font-family:Helvetica; mso-bidi-font-family:Helvetica;color:#2E2E2E;mso-font-kerning:0pt;mso-ansi-language: EN-US'><span style='mso-list:Ignore'>1.<span style='font:7.0pt "Times New Roman"'> </span></span></span><![endif]><span lang=EN-US style='font-size:10.0pt; font-family:"Helvetica",sans-serif;mso-fareast-font-family:"Times New Roman"; mso-bidi-font-family:Arial;color:#2E2E2E;mso-font-kerning:0pt;mso-ansi-language: EN-US'>Dayhoff M, Schwartz R and <span class=SpellE>Orcutt</span> B (1978), <i>"A Model of Evolutionary Change in Proteins"</i>, In Atlas of protein sequence and structure. Vol. 5, pp. 345-352. National Biomedical Research Foundation Silver Spring, MD.<o:p></o:p></span></p>
<p class=MsoListParagraph style='margin-left:18.0pt;mso-para-margin-left:0gd; text-align:justify;text-justify:inter-ideograph;text-indent:-18.0pt;mso-list: l0 level1 lfo1'><![if !supportLists]><span lang=EN-US style='font-size:10.0pt; font-family:"Helvetica",sans-serif;mso-fareast-font-family:Helvetica; mso-bidi-font-family:Helvetica;color:#2E2E2E;mso-font-kerning:0pt;mso-ansi-language: EN-US'><span style='mso-list:Ignore'>2.<span style='font:7.0pt "Times New Roman"'> </span></span></span><![endif]><span class=SpellE><span lang=EN-US style='font-size:10.0pt;font-family:"Helvetica",sans-serif;mso-fareast-font-family: "Times New Roman";mso-bidi-font-family:Arial;color:#2E2E2E;mso-font-kerning: 0pt;mso-ansi-language:EN-US'>Ve</span></span><span lang=EN-US style='font-size: 10.0pt;font-family:"Helvetica",sans-serif;mso-fareast-font-family:"Times New Roman"; mso-bidi-font-family:Arial;color:#2E2E2E;mso-font-kerning:0pt;mso-ansi-language: EN-US'> T, Williams SJ and Kobe B (2015), <i>"Structure and function of Toll/interleukin-1 receptor/resistance protein (TIR) domains"</i>, Apoptosis. Vol. 20(2), pp. 250-261.<o:p></o:p></span></p>
<p class=MsoListParagraph style='margin-left:18.0pt;mso-para-margin-left:0gd; text-align:justify;text-justify:inter-ideograph;text-indent:-18.0pt;mso-list: l0 level1 lfo1'><![if !supportLists]><span lang=EN-US style='font-size:10.0pt; font-family:"Helvetica",sans-serif;mso-fareast-font-family:Helvetica; mso-bidi-font-family:Helvetica;color:#2E2E2E;mso-font-kerning:0pt;mso-ansi-language: EN-US'><span style='mso-list:Ignore'>3.<span style='font:7.0pt "Times New Roman"'> </span></span></span><![endif]><span lang=EN-US style='font-size:10.0pt; font-family:"Helvetica",sans-serif;mso-fareast-font-family:"Times New Roman"; mso-bidi-font-family:Arial;color:#2E2E2E;mso-font-kerning:0pt;mso-ansi-language: EN-US'>Clifton BE and Jackson CJ (2016), <i>"Ancestral Protein Reconstruction Yields Insights into Adaptive Evolution of Binding Specificity in Solute-Binding Proteins"</i>, Cell <span class=SpellE>Chem</span> Biol. Vol. 23(2), pp. 236-245.<o:p></o:p></span></p>
<p class=MsoListParagraph style='margin-left:18.0pt;mso-para-margin-left:0gd; text-align:justify;text-justify:inter-ideograph;text-indent:-18.0pt;mso-list: l0 level1 lfo1'><![if !supportLists]><span lang=EN-US style='font-size:10.0pt; font-family:"Helvetica",sans-serif;mso-fareast-font-family:Helvetica; mso-bidi-font-family:Helvetica;color:#2E2E2E;mso-font-kerning:0pt;mso-ansi-language: EN-US'><span style='mso-list:Ignore'>4.<span style='font:7.0pt "Times New Roman"'> </span></span></span><![endif]><span lang=EN-US style='font-size:10.0pt; font-family:"Helvetica",sans-serif;mso-fareast-font-family:"Times New Roman"; mso-bidi-font-family:Arial;color:#2E2E2E;mso-font-kerning:0pt;mso-ansi-language: EN-US'>Hudson WH, <span class=SpellE>Kossmann</span> BR, de Vera IM, Chuo SW, <span class=SpellE>Weikum</span> ER, <span class=SpellE>Eick</span> GN, Thornton JW, Ivanov IN, <span class=SpellE>Kojetin</span> DJ and <span class=SpellE>Ortlund</span> EA (2016), <i>"Distal substitutions drive divergent DNA specificity among paralogous transcription factors through subdivision of conformational space"</i>, Proc Natl <span class=SpellE>Acad</span> <span class=SpellE>Sci</span> U S A. Vol. 113(2), pp. 326-331.<o:p></o:p></span></p>
<p class=MsoListParagraph style='margin-left:18.0pt;mso-para-margin-left:0gd; text-align:justify;text-justify:inter-ideograph;text-indent:-18.0pt;mso-list: l0 level1 lfo1'><![if !supportLists]><span lang=EN-US style='font-size:10.0pt; font-family:"Helvetica",sans-serif;mso-fareast-font-family:Helvetica; mso-bidi-font-family:Helvetica;color:#2E2E2E;mso-font-kerning:0pt;mso-ansi-language: EN-US'><span style='mso-list:Ignore'>5.<span style='font:7.0pt "Times New Roman"'> </span></span></span><![endif]><span lang=EN-US style='font-size:10.0pt; font-family:"Helvetica",sans-serif;mso-fareast-font-family:"Times New Roman"; mso-bidi-font-family:Arial;color:#2E2E2E;mso-font-kerning:0pt;mso-ansi-language: EN-US'>Wilson C, <span class=SpellE>Agafonov</span> RV, <span class=SpellE>Hoemberger</span> M, <span class=SpellE>Kutter</span> S, Zorba A, <span class=SpellE>Halpin</span> J, <span class=SpellE>Buosi</span> V, <span class=SpellE>Otten</span> R, Waterman D, Theobald DL and Kern D (2015), <i>"Using ancient protein kinases to unravel a modern cancer drug's mechanism"</i>, Science. Vol. 347(6224), pp. 882-886.<o:p></o:p></span></p>
</div>
</body>
</html>